 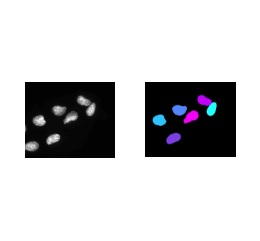
I will be honest IdentifyPrimaryObjects is the second easiest part of the whole segmentation pipeline. You can see the impact of each parameter through the Test Mode feature. I found that the manual parameters in this segmentation module were very easy to work with. So, it is more optimal to select yes and change parameters within the advanced settings (yay… more prototyping…). The only parameters you need to set are the minimum and maximum diameters of the objects, which is easy to do.įollowing the steps in this tutorial video, a user can easily approximate the pixel size of the nuclei in an image.īut, just changing this individual parameter does not work for every dataset. When first applying the pipeline, this module does not show the “advanced settings”, which makes it more approachable. Starting with the IdentifyPrimaryObjects module, our goal is to identify the nuclei in the DAPI or Hoechst images. IdentifyTertiaryObjects: Segment cytoplasm by subtracting the nuclei from the whole cell outline IdentifySecondaryObjects: Segment whole cells using an RNA channel IdentifyPrimaryObjects: Segment nuclei from a DAPI/Hoechst channel The standard segmentation pipeline for Cell Painting images with CellProfiler is as follows: CellProfilerĬellProfiler might have been difficult to work with for illumination correction, but CellProfiler excels in segmentation. Note: My segmentation findings in this blog are based on Cell Painting data.įor more information on Cell Painting assays and what makes them unique, see the following GitHub wiki from the Broad Institute. I will discuss these methods in part II of my segmentation blog. I also tested Ilastik, Weka Trainable Segmentation, and Scikit testing.įrom my experience, these performed sub-optimally with my data. I will be comparing multiple segmentation methods in this blog post, which are: I recommend benchmarking different methods with your own data. If you are just starting with this blog post, then welcome in!Īs a reminder from my previous post, keep in mind that any method or software you use in your pipeline might not be optimal or might not contain the functionalities that you want (e.g. Welcome back! I hope that you were able to come up with an amazing illumination correction pipeline from my illumination correction blog post and are ready for segmentation! A Brief Comparison of Popular Cell Segmentation Software 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
